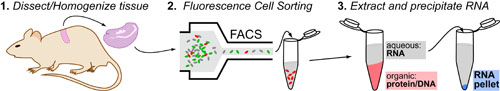
Quantitiative Reverse-Transcriptase PCR (RT-PCR) is different from regular PCR in that it does not measure genes(i.e. DNA) per se but rather measures the EXPRESSION of those gene (i.e. mRNA). Given the fact gene expression (mRNA) can vary dramatically between cell-types, it is important to first isolate a single cell-type by:
- Dissect and homogenize organ/tissue containing cells of interest
- Purify/Sort cell-type using a technique like Flow Cytometry
- Extract RNA using Guanidiunum thiocyanate(denatures proteins) and phenol-chloroform (removes protein/DNA)
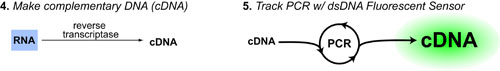
Once you have purified the RNA from your cells you must use reverse transcriptase to transform that RNA into cDNA (complementary DNA). Finally it is the cDNA that you PCR amplify. In addition, fluorescent sensors which detect dsDNA are used to monitor the time course of the PCR yeilding the data pictured below:
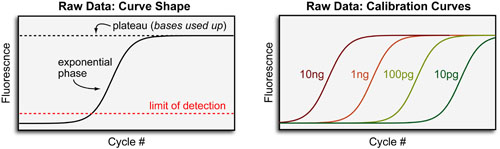
In general, quantitative PCR (for measuring DNA) and RT-PCR (for measuring RNA) data has a sigmoidal (“s-like”) shape featuring a pleateau at high cycle numbers when the primers/bases are used up. As can be seen in a model calibration curve below: higher mRNA expression is indicated by left-ward shift of the curve. Given the varying PCR efficiencies between different primers/genes, absolute quantification of mRNA requires calibration curves for every gene/experiment. If only relative quantification of mRNA is required, it is typically sufficient to simultaneously monitor a “reference” or “house-keeping” gene that is known to be stably expressed at similar levels between different cells.
REFERENCES:
- Chomczynski, P.; Sacchi, N. Single-Step Method of RNA Isolation by Acid Guanidinium Thiocyanate-Phenol-Chloroform Extraction Anal. Biochem., 1987
- Bustin, S.A. Why the need for qPCR publication guidlines? – The case for MIQE. Methods, 2010, 217-226.
- Schefe, J.H.; Lehmann, K.E.; Buschmann, I.R.; Unger, T.; Funke-Kaiser, H. Quantitative real-time RT-PCR data analysis: current concepts and the novel “gene expressions’s CT difference” formula. J.Mol. Med. 2006 DOI 10.1007/s00109-006-0097-6
- Schmittgen, T.D.; Livak, K.J. Analyzing real-time PCR data by the comparative CT method. Nat. Protocols, 2008, 3, 1101-1108.
- Molecular Cloning: Technical Guide v. 2, 2014, New England Biolabs Inc.
- Alberts, B. Molecular Biology of the Cell 5th Ed. Garland Science 2008
- Murphy, K. Janeway’s Immunobiology 8th Ed. Garland Science 2012

This work by Eugene Douglass and Chad Miller is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License.